Inbred mice

2
Biostatistics & Medical Informatics, University of Wisconsin – Madison

2

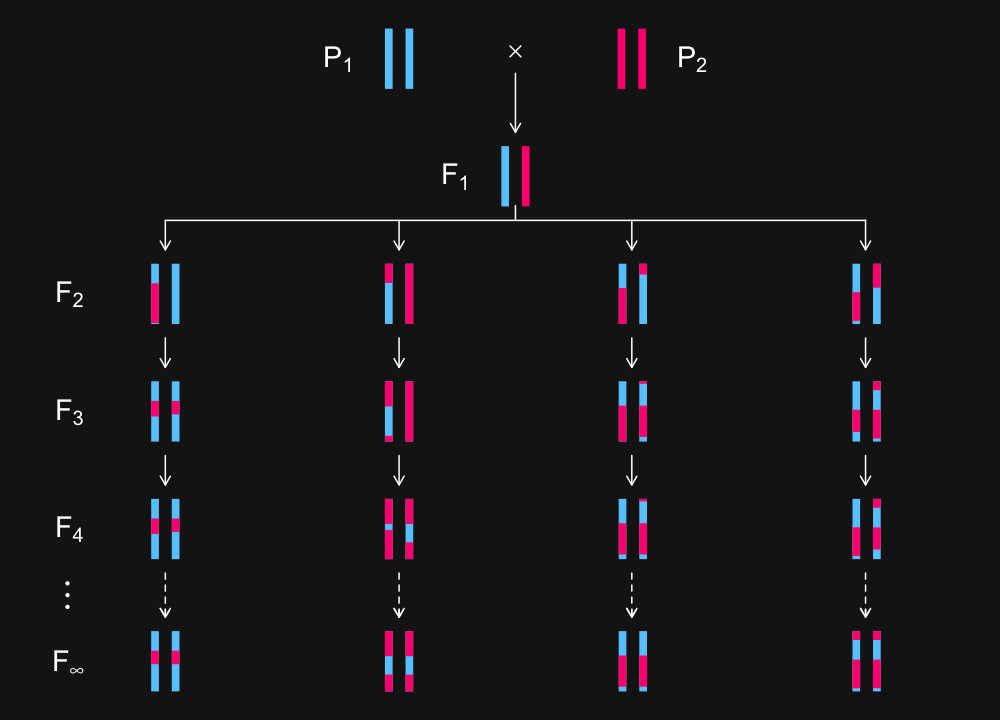
3
4
5
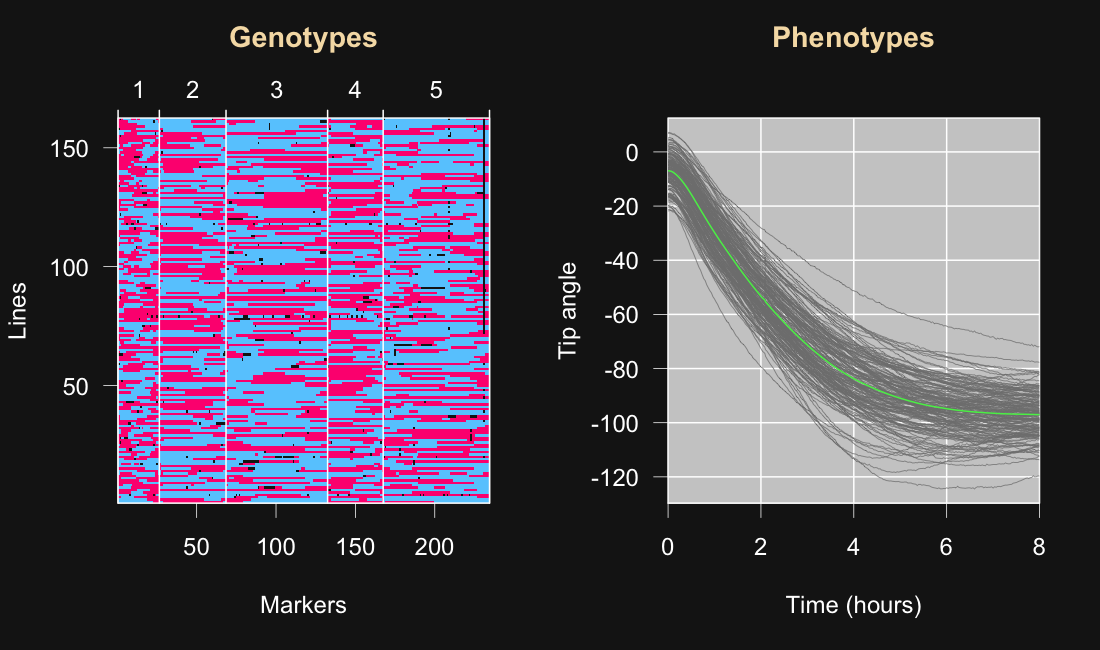
6
7
8
9
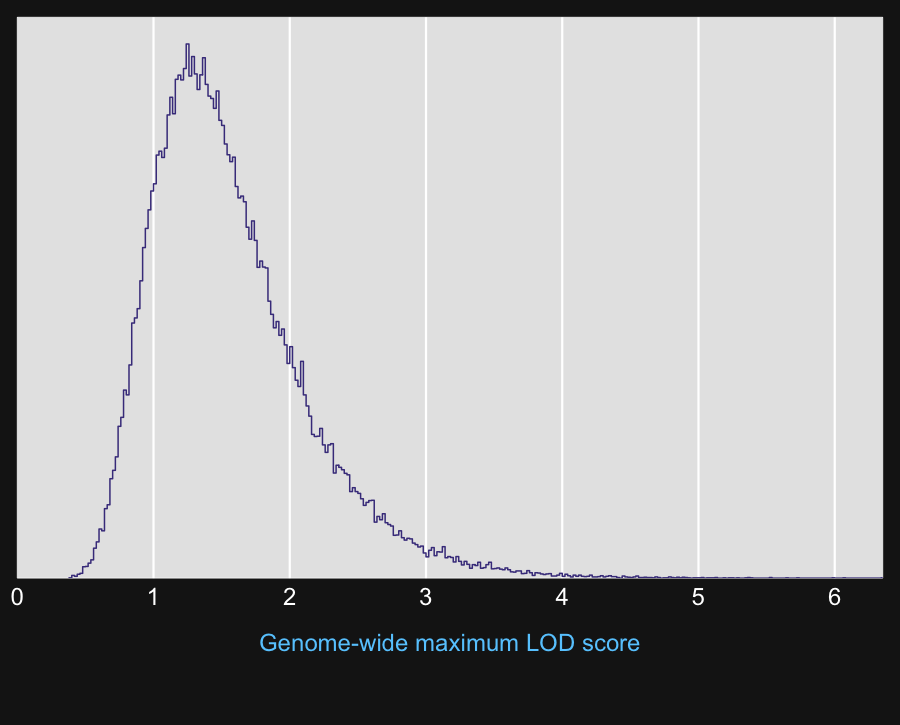
10
11
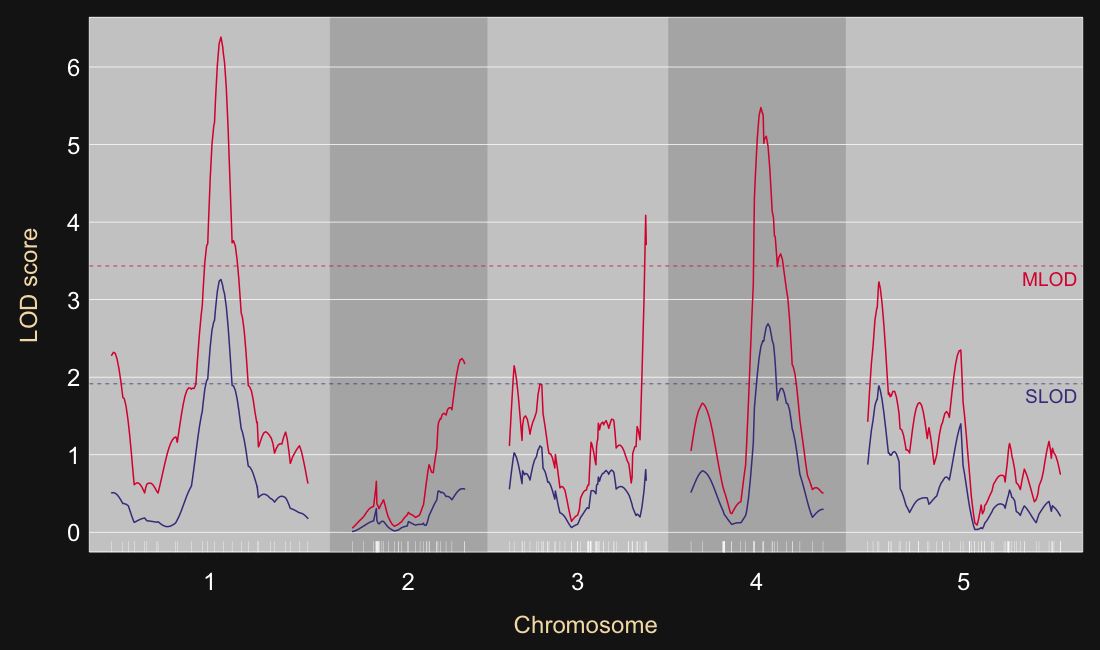
SLOD(λ) = avet LODt(λ)
MLOD(λ) = maxt LODt(λ)
12
13
14
Among additive QTL models, maximize
pLOD(γ) = LOD(γ) – T × |γ|
γ = a multiple-QTL model
|γ| = no. QTL in the model
T = significance threshold
KW Broman, TP Speed (2002) J Roy Stat Soc B 64:641-656
15
Among additive QTL models, maximize
pLODt(γ) = LODt(γ) – T × |γ|
γ = a multiple-QTL model
|γ| = no. QTL in the model
T = significance threshold
KW Broman, TP Speed (2002) J Roy Stat Soc B 64:641-656
15
16
16
16
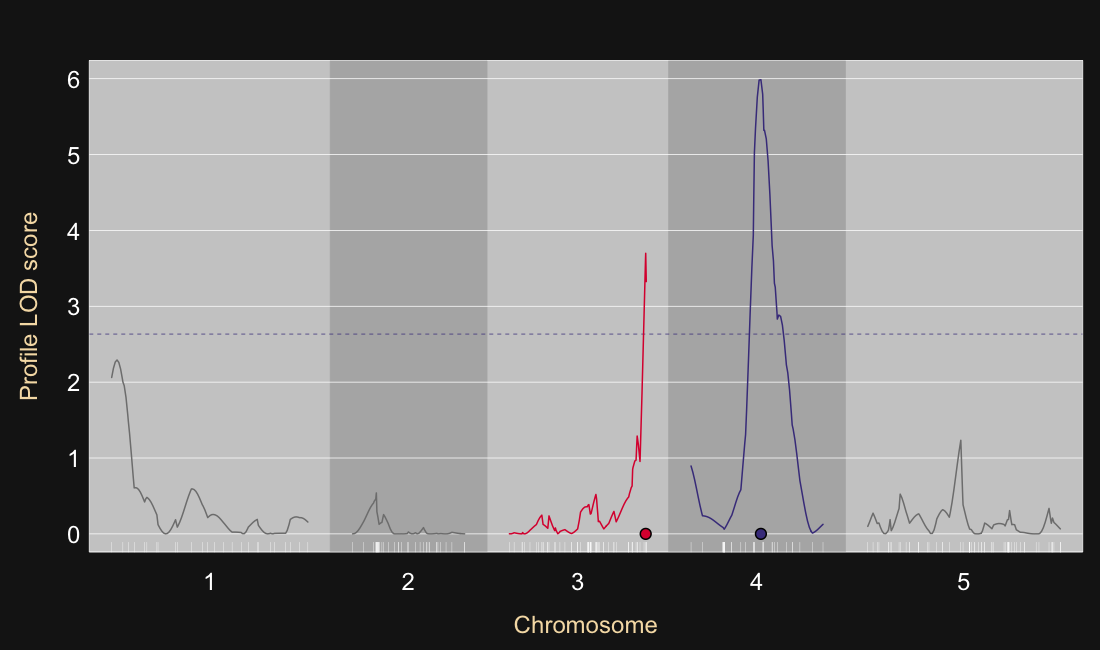
17
18
Among additive QTL models, maximize
pSLOD(γ) = avet LODt(γ) – TS × |γ|
pMLOD(γ) = maxt LODt(γ) – TM × |γ|
γ = a multiple-QTL model
|γ| = no. QTL in the model
T = penalties
19

20
21
| Edgar Spalding | Botany, UW-Madison | |
| Candace Moore | ||
| Logan Johnson | ||
| Il-youp Kwak | Statistics, UW-Madison | |
| NIH: R01 GM074244 | ||
22