|
||
|
Yuan and Kendziorski (2006) proposed an approach to identify
differentially expressed (DE) genes across multiple biological conditions over time.
The approach provides an FDR controlled list of
interesting genes that are DE at each time point, and over time.
Details of the approach can be found in Yuan and Kendziorski,
Journal of the American Statistical Association 101(476): 1323-1332; With discussion 1332-1340.
The EB-HMM package can be downloaded here,
which contains the R code for running the EB-HMM approach along with a demo. If that does not work (there seems to be a problem with some browsers), you can access the tar gzipped file
here.
To run the demo, install the package in a directory, start R, and use
the commands below:
library("HMM, lib.loc=".") # if R is started from the directory in which the
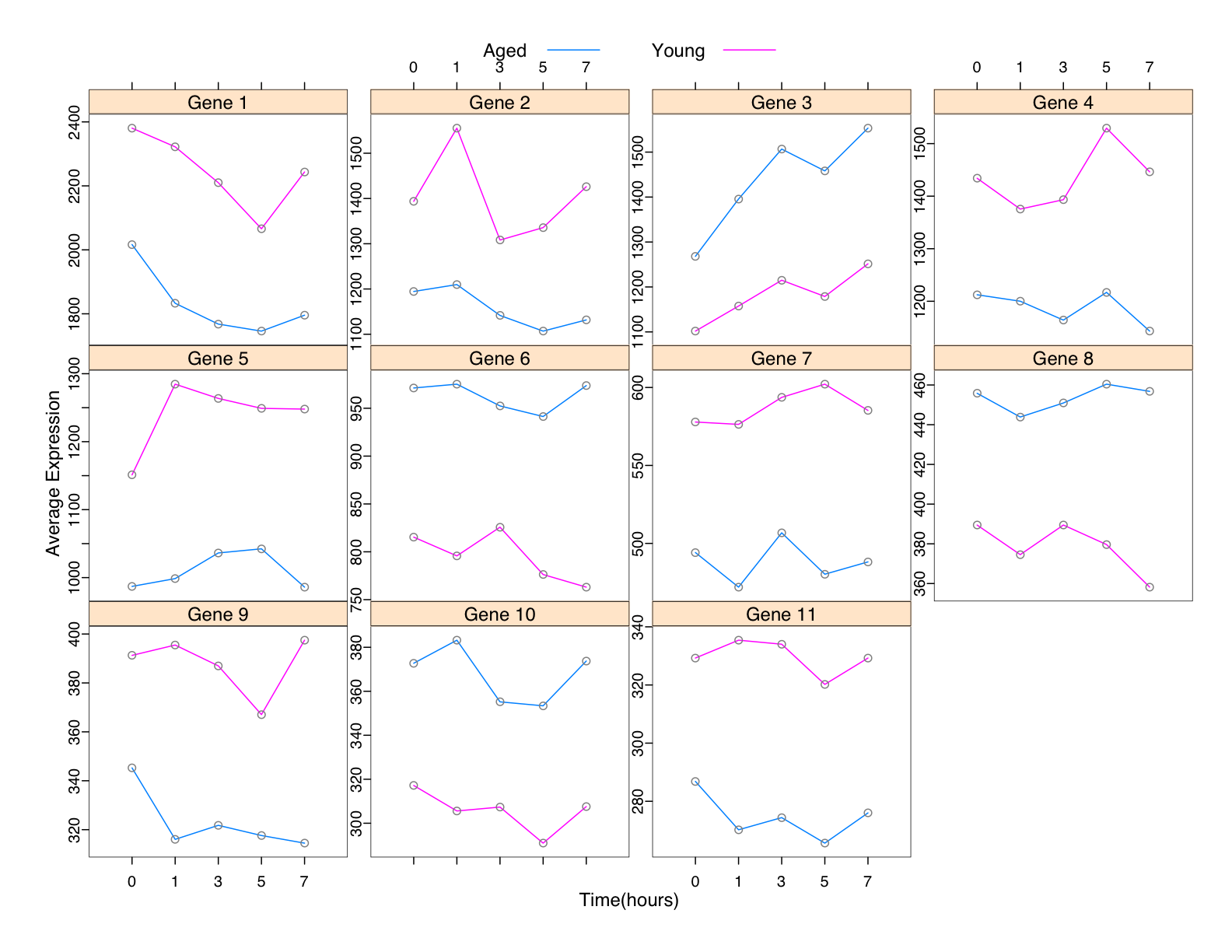
package is installed; otherwise specify the location. Posterior probabilities are provided for each expression pattern at each time. In the majority of case studies we've considered, the approach proves to be more powerful at identifying genes over time than approaches that consider each time point in isolation. This is because EB-HMM shares information across time points, and so genes such as those shown in Figure 1 may be identified. In particular, we highlight 11 genes identified in a case study detailed in Yuan and Kendziorski, JASA, 2006. Each of the 11 genes was declared EE at every time point by a number of methods, but DE at every time point by EB-HMM. As shown, the differences in average expression are relatively small, but the changes are consistent over time, and as a result our EB-HMM approach finds them.
Figure 1: Shown are 11 genes from an experiment detailed in Yuan and Kendziorski, JASA, 2006. The pink (blue) corresponds to expression from young (old) animals in a study of oxidative stress on the heart and how it changes with age. Each point at each time represents expression averaged across 3 animals. Last Modified June 2009 |