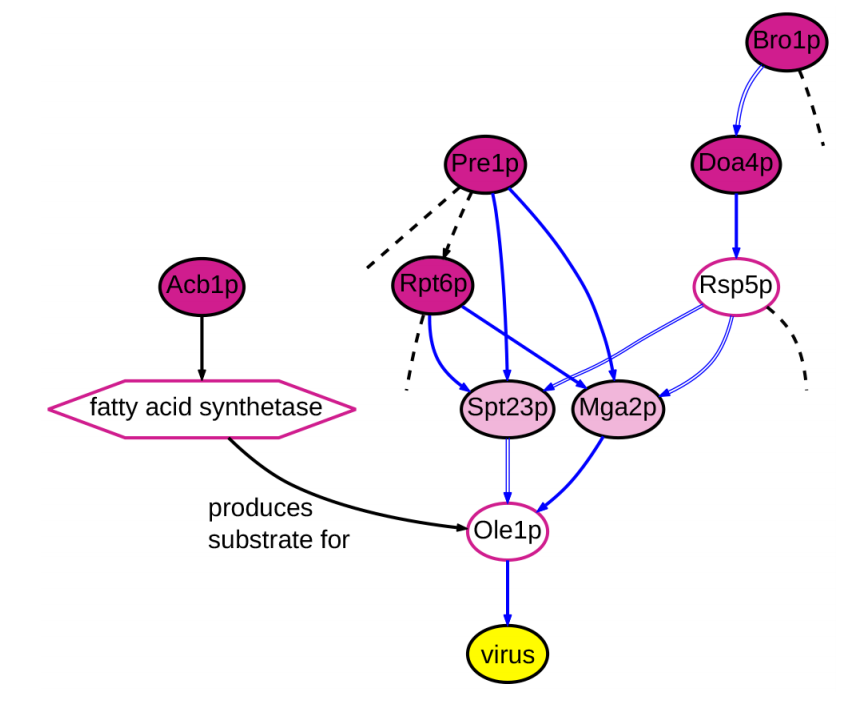
Current projects in my group include the following:
Characterizing host virus interactions including the host networks involved in viral replication, and viral-genotype to disease-phenotype associations.
Representative publications:L. Hao, Q. He, Z. Wang, M. Craven, M. Newton and P. Ahlquist (2013).
Limited Agreement of Independent RNAi Screens for Virus-Required Host Genes Owes More to False-Negative than False-Positive Factors.
PLoS Computational Biology 9(9).
K. Lee, A. Kolb, Y. Sverchkov, J. Cuellar, M. Craven and C. Brandt (2015).
Recombination Analysis of Herpes Simplex Virus Type 1 Reveals a Bias towards GC Content and the Inverted Repeat Region.
Journal of Virology.
D. Chasman, B. Gancarz, L. Hao, M. Ferris, P. Ahlquist and M. Craven (2014).
Inferring Host Gene Subnetworks Involved in Viral Replication.
PLoS Computational Biology 10(5).
Inferring intracellular networks from genome-wide experiments.
Representative publications:D. Chasman, Y.-H. Ho, D. Berry, C. Nemec, M. MacGilvray, A. Merrill, J. Hose, M. V. Lee, J. Will, J. Coon, A. Ansari, M. Craven and A. Gasch (2014).
Pathway Connectivity and Signaling Coordination in the Yeast Stress-Activated Signaling Network.
Molecular Systems Biology 10(11):759.
S. Kiblawi D. Chasman, A. Henning, E. Park, H. Poon, M. Gould, P. Ahlquist and M. Craven (2019).
Augmenting subnetwork inference with information extracted from the scientific literature.
PloS Computational Biology 15(6):e1006758.
Learning models to assess risk for clinical events such as asthma exacerbations and post-hospitalization VTEs from electronic health records.
Representative publications:S. I. Feld, A. G. Cobian, S. E. Tevis, G. D. Kennedy and M. W. Craven (2016).
Modeling the Temporal Evolution of Postoperative Complications
Proceedings of the American Medical Informatics Association (AMIA) Annual Symposium.
E. Kawaler, A. Cobian, P. Peissig, D. Cross, S. Yale and M. Craven (2012).
Learning to Predict Post-Hospitalization VTE Risk from EHR
Data.
Proceedings of the American Medical Informatics
Association (AMIA) Annual Symposium.
Extracting information from the scientific literature and exploiting this information for understanding biological data.
Representative publications:H. Shatkay and M. Craven (2012).
Mining the Biomedical Literature.
MIT Press.
A. Vlachos & M. Craven (2012).
Biomedical Event Extraction from Abstracts and Full Papers using Search-Based Structured Prediction.
BMC Bioinformatics 13(Suppl. 11):S5.